Theoretical Biophysics
Our multidisciplinary research group studies dynamical systems in biology. We address how macromolecular self-assembly is controlled to occur at the right place and time, with focus on endocytosis, viral exit, and transcriptional regulation. Group members are interested both in principles and design of self-assembly, as well as biological function in cells. They develop and apply theory, modeling, and simulations to these problems, with frequent collaboration with experimentalists.
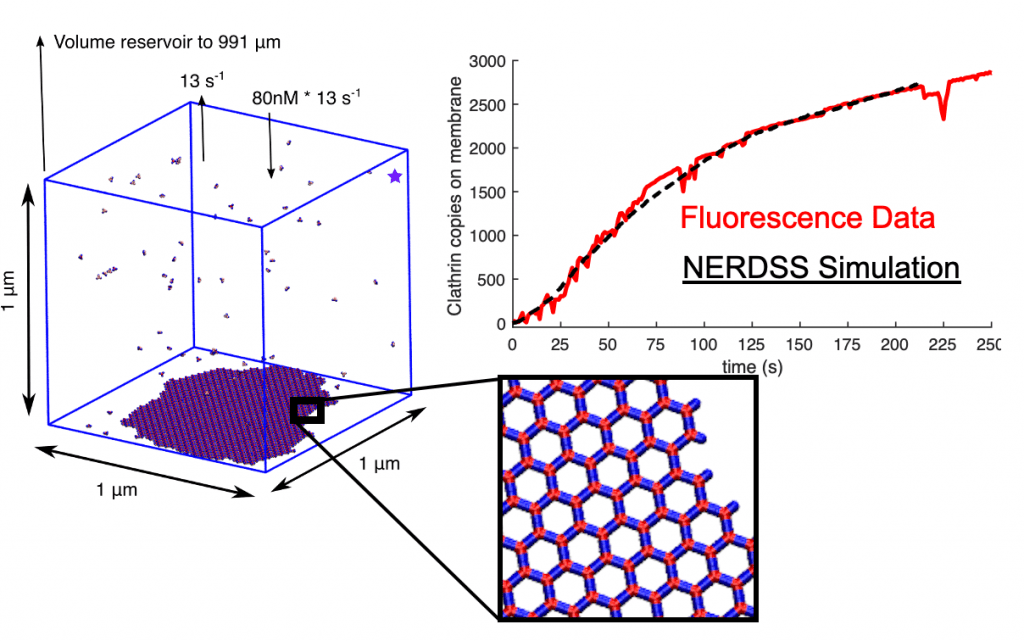
*Positions available for post-docs and grad students! Projects related to membrane remodeling by protein assemblies, virus/HIV budding and assembly, and temporal regulation of DNA transcription/repair*
twitter/X @mejohnson81
News and Announcements
June 2024 Welcome new PhD student Smriti Chhibber and new Master’s student Ruihan Quan to the Johnson Lab for graduate research!
June 2024 The Johnson lab research is highlighted in JHU’s Arts and Sciences Magazine
March 2024 Check out Maggie’s “Living Histories” video on youtube, a series where faculty share their path to becoming scientists and professors in 10-minute episodes.
Nov 2023 Good-bye to post-doc Dr. Yiben Fu, congrats on starting your faculty position at South China University of Technology!
July 2023 Welcome new post-doc Sam Foley!
June 2023 Congrats to graduating senior Yian Qian, our latest superstar first-author undergrad to head off to graduate school!
June 2022 Congrats to Maggie on receiving the official tenure decision and promotion to Associate Professor at Johns Hopkins!
May 2022 Congrats to graduating seniors Jackie Chang and Nandan Kulkarni, good luck Nandan in your PhD research next year!
Oct 2021 Welcome Master’s student Zixiu Hugh Liu to the Johnson Lab for your PhD research!
Sept 2021 Welcome graduate students Jonathan Fischer from Biology and Moon Ying from Chem BE to the Johnson Lab for your PhD research!
June 2021 Welcome Summer REU student Millie Lewis!
May 2021 Welcome Mankun Sang to the Johnson Lab for your PhD research!
May 2021 Welcome Adip as an official PhD student in the Johnson Lab!
May 2020 Congrats Adip Jhaveri on completing a Master’s at JHU, welcome back for your PhD!
Jan 2020 Welcome new post-doc Dr. Sikao Guo!
August 2019 Good luck to recent grad Daisy Duan in the Biophysics PhD program at Yale!
July 2019 Welcome new post-doc Bhavya Mishra!
July 2019 Congratulations to Maggie for being awarded an NIH MIRA!
July 2018 Welcome new post-doc Yi-Ben Fu!
June 2018 Welcome BRBT summer high-school student Taqeeyah to the group!
June 2018 Welcome Yasmin Moghadamnia as a new graduate student to the group!
May 2018 Congratulations to Maggie for winning an NSF CAREER Award from the Chemistry division!
Dec 2017 Congratulations to Dr. David Holland for defending his dissertation and receiving his PhD in Biomedical Engineering!
