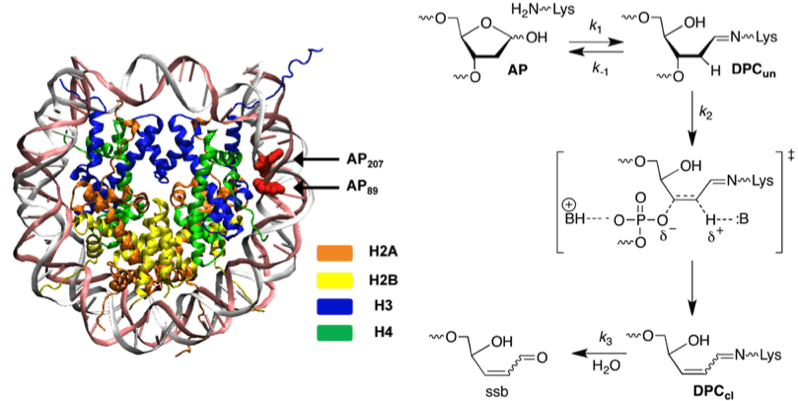
DNA lesions are biologically deleterious because they can adversely affect DNA replication and transcription. Little attention has been paid to the possibility that lesions may react within the biopolymer to produce in more deleterious forms of damage. Furthermore, most studies have been carried out on naked DNA. We have shown that histone proteins in nucleosome core particles catalyze the cleavage of alkali labile lesions, such as abasic sites (AP). The histone proteins accelerate strand scission as much as 1500-fold. Abasic site reactivity was investigated using organic synthesis, kinetics, mass spectrometry, and site-specific mutagenesis.
The lysine rich histone proteins catalyze the cleavage at (oxidized) abasic sites via a lyase mechanism reminiscent of DNA repair enzymes (above). Interestingly, the aspartate, glutamate, and histidine content of histone tails are significantly lower than in proteins in general. While the carboxylate amino acids are likely absent to help control transcription, our experiments suggest that histone proteins may have evolved to minimize histidine content so as to protect damaged DNA from undergoing strand scission.
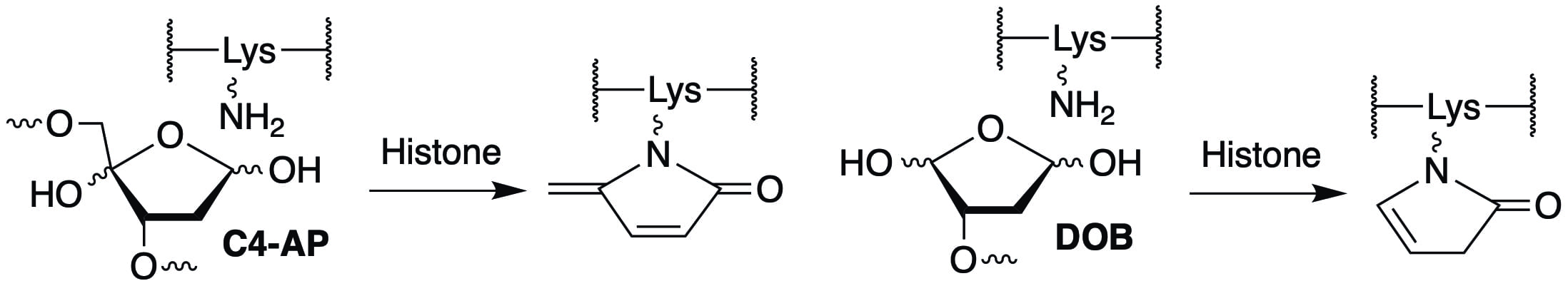
The lysine rich amino terminal tails of the histone proteins, which are often modified post-translationally in cells are primarily responsible for catalyzing strand scission. In addition, histone mediated cleavage of oxidized abasic lesions (C4-AP, DOB) produced by several antitumor agents that damage DNA also results in modification of the lysine rich tail. These modifications raise new questions concerning the possibility that they act as post-translational modifications and affect the histone proteins’ interaction with proteins involved in genetic regulation. We are actively pursuing these questions.
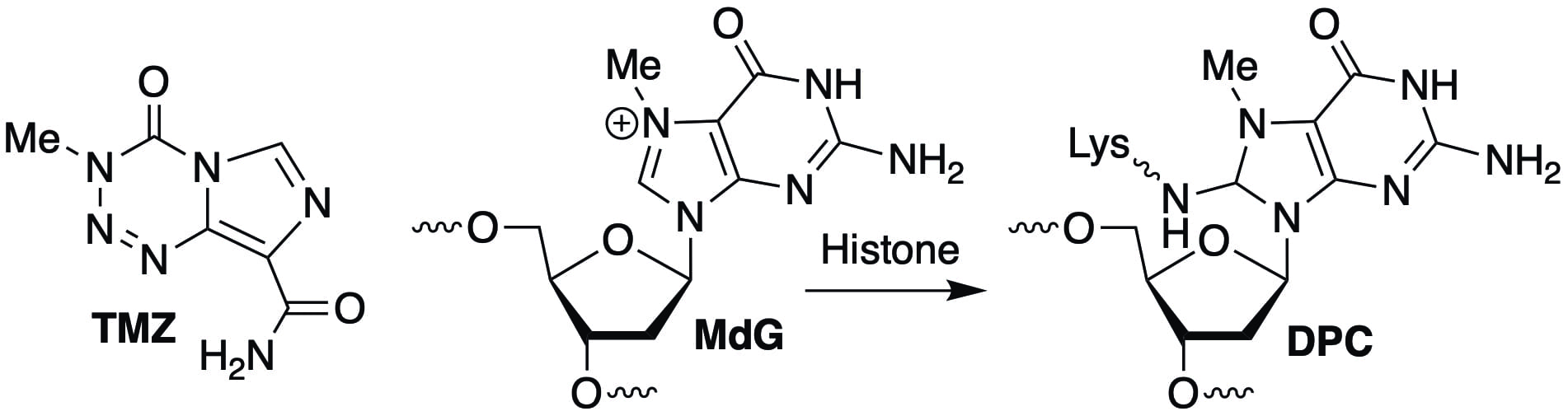
Recently, we discovered that N7-methyl-2′-deoxyguanosine (MdG), the major DNA lesion produced by antitumor agents such as temozolomide (TMZ; used to treat glioblastoma) forms DNA-protein cross-links (DPCs) in cells. Studies on nucleosome core particles showed that the DPCs result from reaction of the lysine residues within histone proteins. This fundamental observation could alter how cancer cell biologists think about how monofunctional DNA alkylating agents such as TMZ kill cells.
For relevant publications see:
• Rapid DNA-Protein Cross-linking and Strand Scission by an Abasic Site in a Nucleosome Core Particle. Sczepanski, J. T.; Wong, R. S.; McKnight, J. N.; Bowman, G. D.; Greenberg, M. M. Proc. Natl. Acad. Sci. USA 2010, 107, 22475-22480.
• Histone Catalyzed Cleavage of Nucleosomal DNA Containing 2-Deoxyribonolactone. Zhou, C.; Greenberg, M. M. J. Am. Chem. Soc. 2012, 134, 8090-8093. “Spotlighted” in the J. Am. Chem. Soc. 2012, 134, 9031.
• Mechanistic Studies on Histone Catalyzed Cleavage of Apyrimidinic/Apurinic Sites in Nucleosome Core Particles. Zhou, C.; Sczepanski, J. T.; Greenberg, M. M. J. Am. Chem. Soc. 2012, 134, 16734-16741.
• Histone Modification via Rapid Cleavage of C4′-Oxidized Abasic Sites in Nucleosome Core Particles. Zhou, C.; Sczepanski, J. T.; Greenberg, M. M. J. Am. Chem. Soc. 2013, 135, 5274-5277. “Spotlighted” in the J. Am. Chem. Soc. 2013, 135, 5933.
• Rapid Histone Catalyzed DNA Lesion Excision and Accompanying Protein Modification in Nucleosomes and Nucleosome Core Particles. Weng, L.; Greenberg, M. M. J. Am. Chem. Soc. 2015, 137, 11022-11031.
• Histone Tail Sequences Balance Their Role in Genetic Regulation and the Need to Protect DNA Against Destruction in Nucleosome Core Particles Containing Abasic Sites. Yang, K.; Greenberg, M. M. ChemBioChem (accepted).
• Histone Tails Decrease N7-Methyl-2′-Deoxyguanosine Depurination and Yield DNA-Protein Crosslinks in Nucleosome Core Particles and Cells. Yang, K.; Park, D.; Tretyakova, N. Y.; Greenberg, M. M. Proc. Natl. Acad. Sci. USA 2018, 115, E11212-E11220. Highlighted in Chemical & Engineering News (https://cen.acs.org/biological-chemistry/nucleic-acids/Chemicals-cause-unexpected-DNA-damage/96/i48)